 |
ScaLAPACK 2.1
2.1
ScaLAPACK: Scalable Linear Algebra PACKage
|
 |
ScaLAPACK 2.1
2.1
ScaLAPACK: Scalable Linear Algebra PACKage
|
#include "../pblas.h"#include "../PBpblas.h"#include "../PBtools.h"#include "../PBblacs.h"#include "../PBblas.h"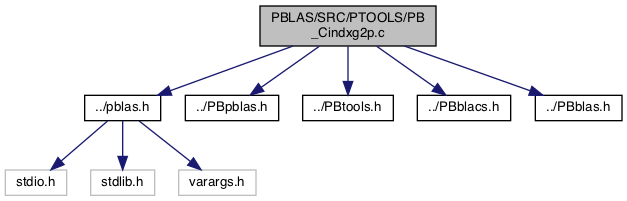
Go to the source code of this file.
Functions | |
| int | PB_Cindxg2p (int IG, int INB, int NB, int PROC, int SRCPROC, int NPROCS) |
| int PB_Cindxg2p | ( | int | IG, |
| int | INB, | ||
| int | NB, | ||
| int | PROC, | ||
| int | SRCPROC, | ||
| int | NPROCS | ||
| ) |
Definition at line 22 of file PB_Cindxg2p.c.